LipidBlast-mz-lookup - explains how to perform an MS1 search only and discusses the needed resolving power LipidBlast-Templates - this explains how to study and create lipid fragmentations using EXCEL Lipid-Classes -this link details about the covered classes and number of spectra in LipidBlast Our LipidBlast library works with low-resolution and high-resolution instruments. 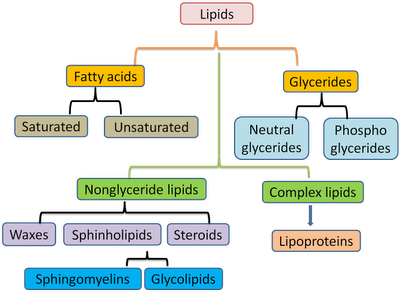
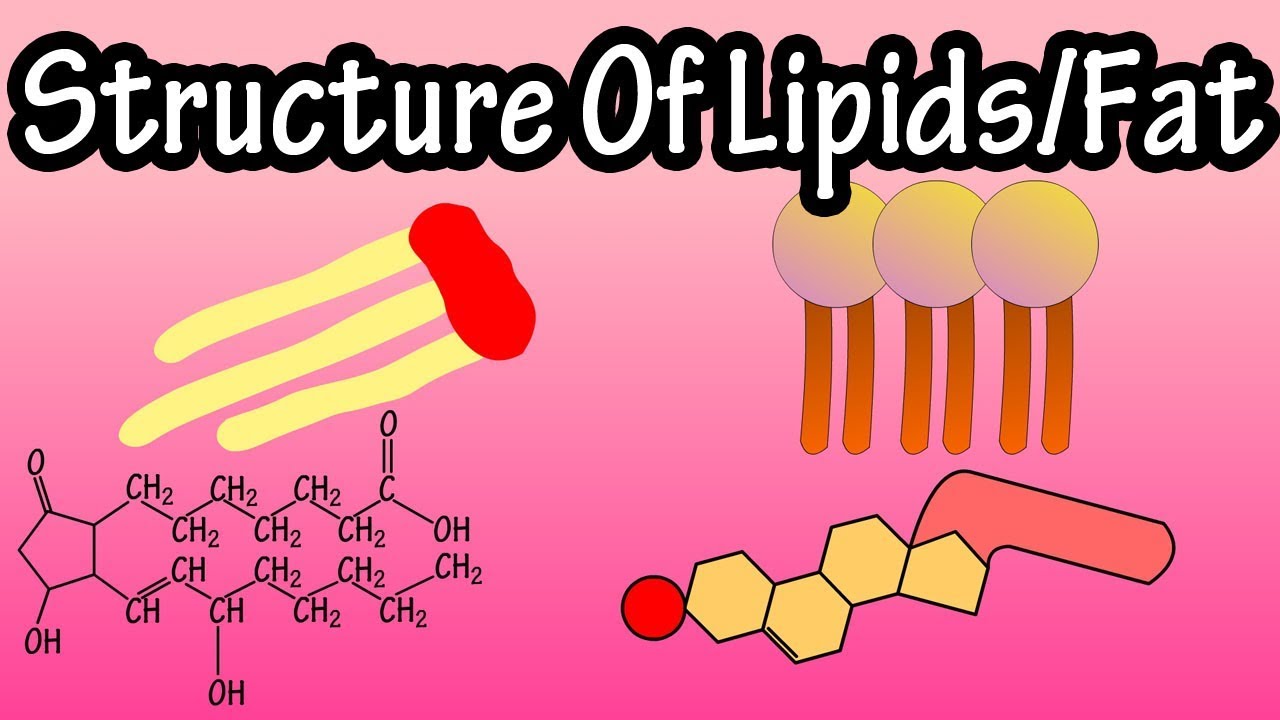
We developed a computer-generated (in-silico) tandem mass spectral (MS/MS) database that can be used to annotate and identify hundreds of lipids in plants, bacteria, algae, animals, humans, viruses. Lipid analysis of polar lipids can be performed with tandem mass spectrometry and mass spectral library search. PubMed Central Link (Open Access): PMC3731409 Published in Nature Methods (2013) Advanced Online LINK: doi:10.1038/nmeth.2551 Results: LipidBlast in silico tandem mass spectrometry database for lipid identification Project Partner: Tobias Kind, Kwang-Hyeon Liu, Do Yup Lee, Brian DeFelice, John K.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
